FusionSeq Gallery
From GersteinInfo
User documentation main
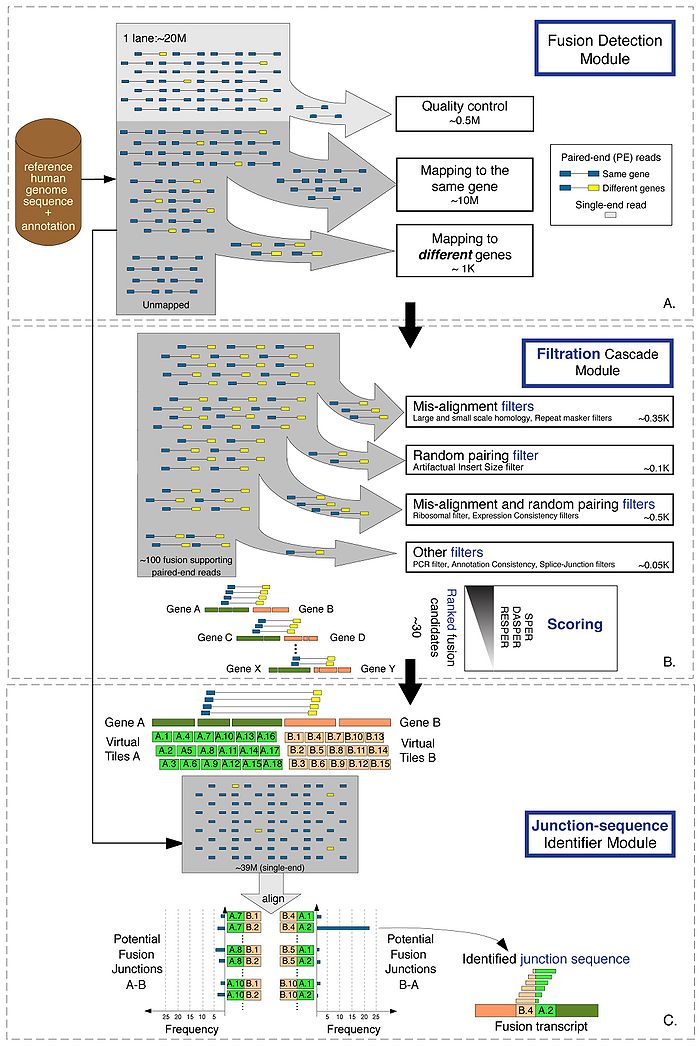
Schematic of FusionSeq.
- A. The PE reads are processed to identify potential fusion candidates. Poor quality reads are discarded at first, and the remaining PE reads are aligned to the reference human genome (hg18). The reads are compared to the annotation set (UCSC knownGenes) in order to classify them as belonging to the same gene or to different genes. Those aligned to two different genes are then selected as potential fusion candidates. All good quality single- end reads are also stored for the identification of the sequence of the junction.
- B. The filtration cascade module analyzes the candidates and removes those that have high sequence homology between the two genes or a higher insert-size compared to the transcriptome norm. Additional filters are employed to remove candidates due to random pairing and misalignment as well as PCR artifacts and annotation inconsistencies. The high-confidence list of candidates is then scored and processed to find the sequence of the junction.
- C. The junction-sequence identifier detects the actual sequence at the breakpoints by constructing a fusion junction library. It first covers the regions of the potential breakpoint of each gene with “tiles” 1bp apart, and then creates all possible combinations, considering both orientation of the fusion, namely gene A upstream of gene B and vice versa. All single-end reads are then aligned to the fusion junction library and the junction with the highest support is identified as the sequence of the fusion transcript junction.